Publications
- 1
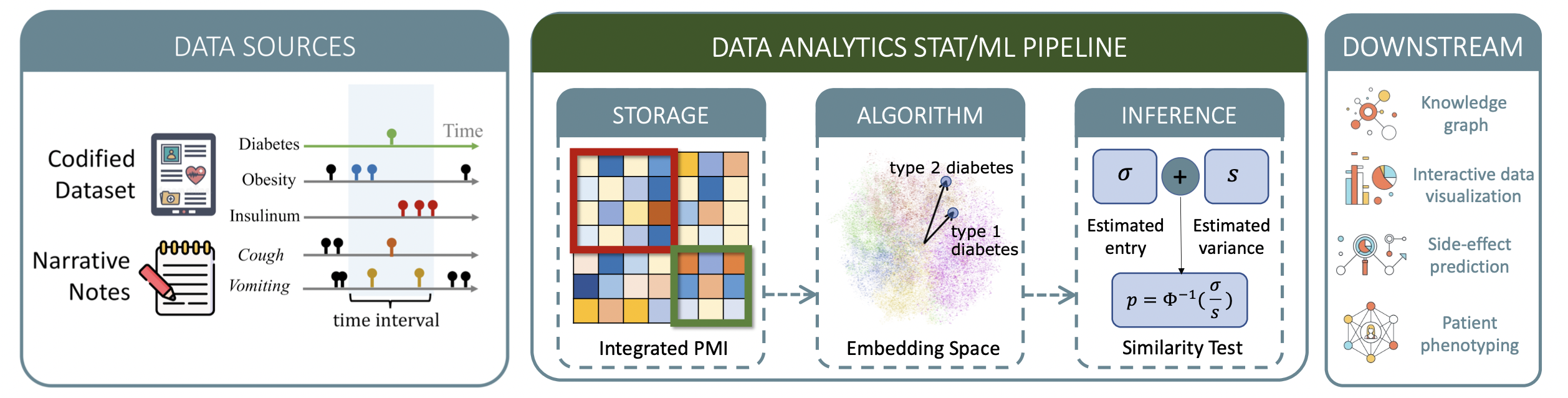
ARCH: Large-scale Knowledge Graph via Aggregated Narrative Codified Health Records Analysis
The ARCH algorithm addresses codified-data and clinical-note integration by constructing a large-scale knowledge graph with over 60,000 clinical concepts from 12.5 million Veterans Affairs patients. It derives embedding vectors from co-occurrence matrices, quantifies relatedness with cosine similarity and p-values, and applies sparse embedding regression to remove indirect linkages with statistical certainty. Validation across disease phenotyping, drug side-effect prediction, and Alzheimer’s subtyping shows robust performance beyond pure text models such as PubMedBERT.
- 2
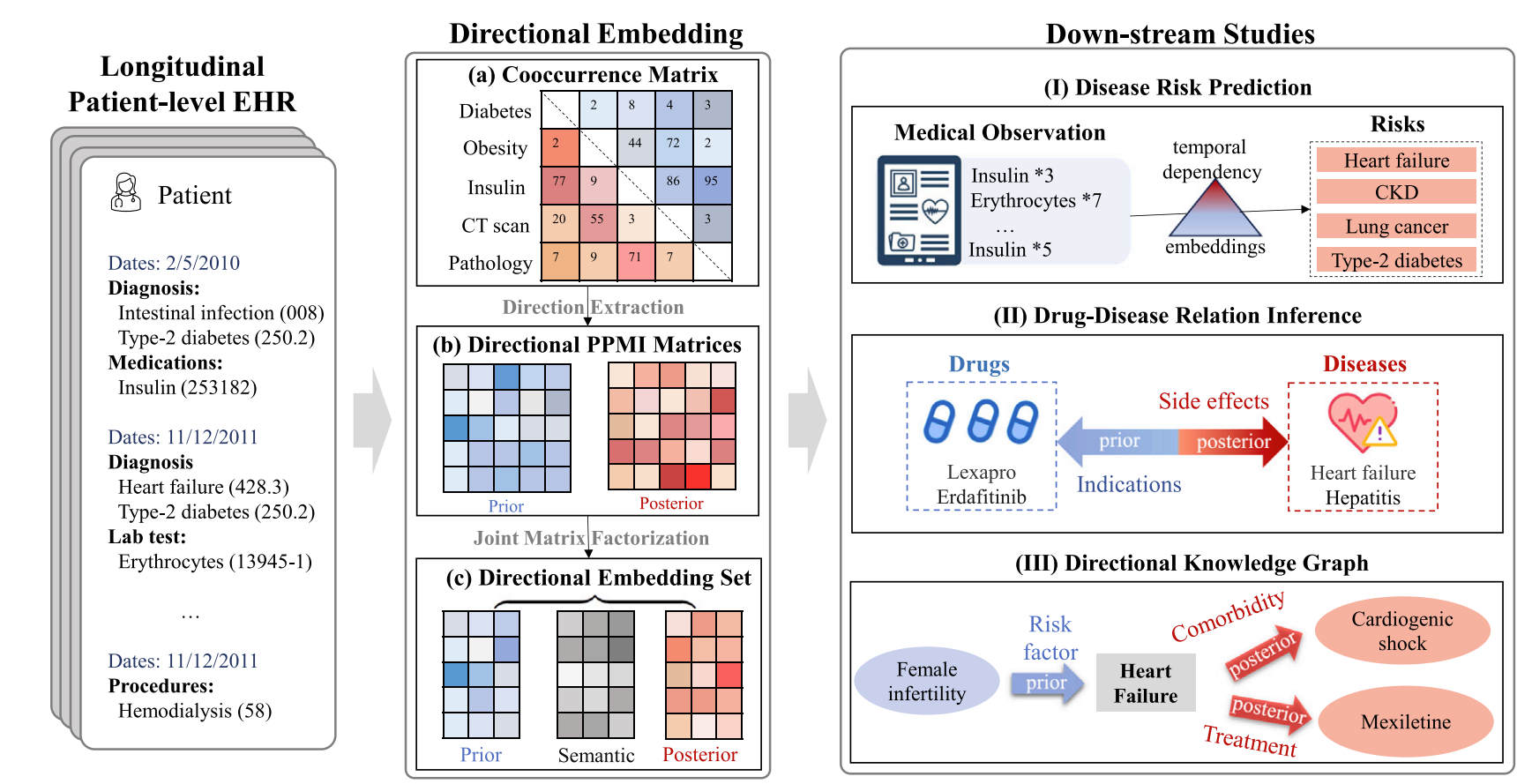
DOME: Directional medical embedding vectors from Electronic Health Records
DOME captures temporally directional dependencies between clinical concepts using summary-level EHR data. It aggregates patient data into asymmetric co-occurrence matrices, computes dual PPMI matrices, and outputs shared semantic, prior-context, and posterior-context embeddings for each concept. Validation studies show improved risk prediction, unsupervised distinction between drug indications and side effects, and directional knowledge-graph construction while preserving patient privacy.
- 3
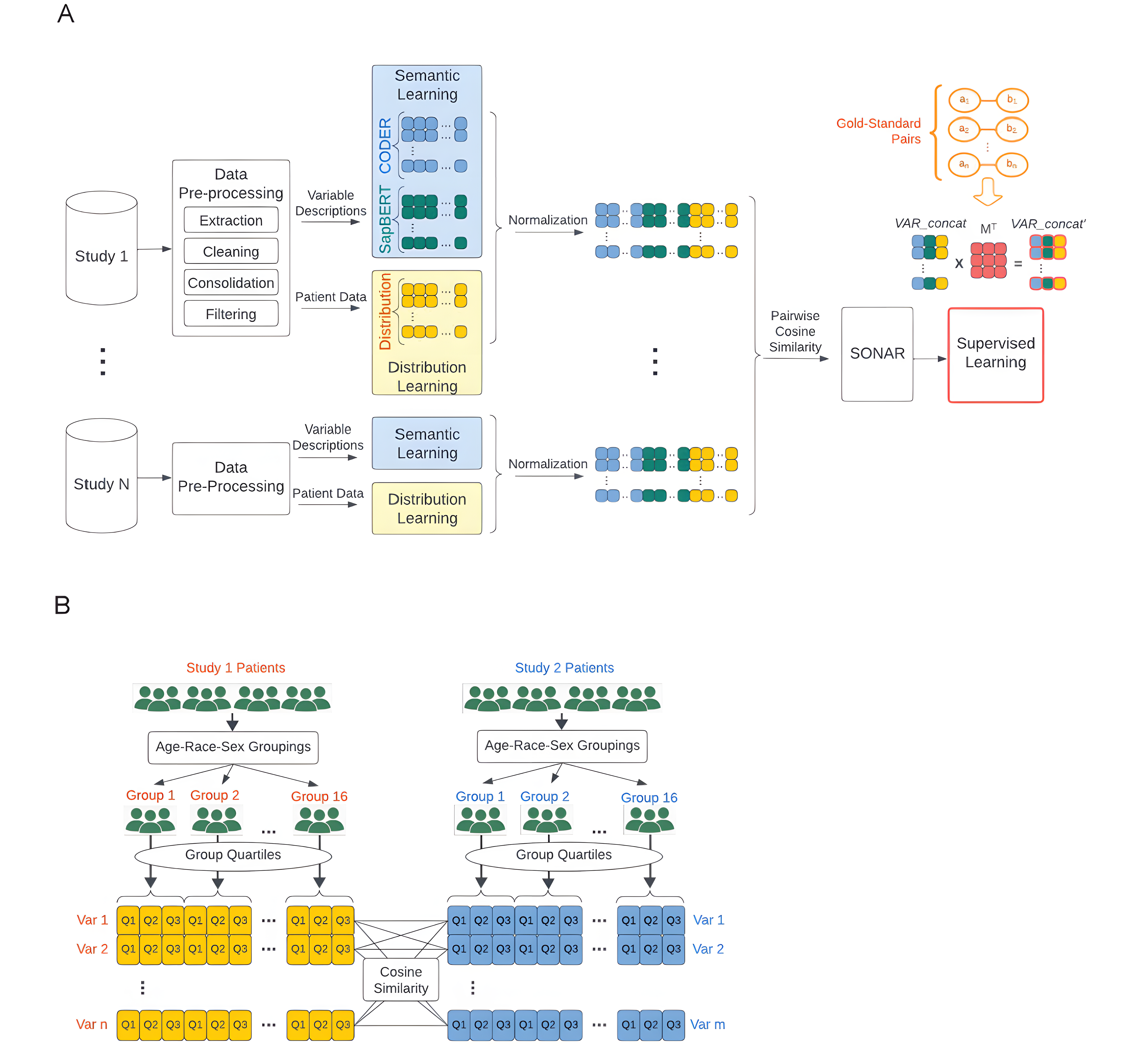
Robust automated harmonization of heterogeneous data through ensemble machine learning: algorithm development and validation study
SONAR combines semantic learning from variable descriptions with distribution learning from cohort data to solve variable misalignment in multi-cohort studies. It leverages CODER and SapBERT features together with subgroup distribution statistics, including quartile-based vectors across demographic subgroups, to map local concepts to standardized vocabularies. Supervised SONAR outperforms semantic-only approaches, especially for hard concepts with weak textual descriptions, reducing manual curation burden for large-scale integrative research.
- 4
Representation learning to advance multi-institutional studies with electronic health record data
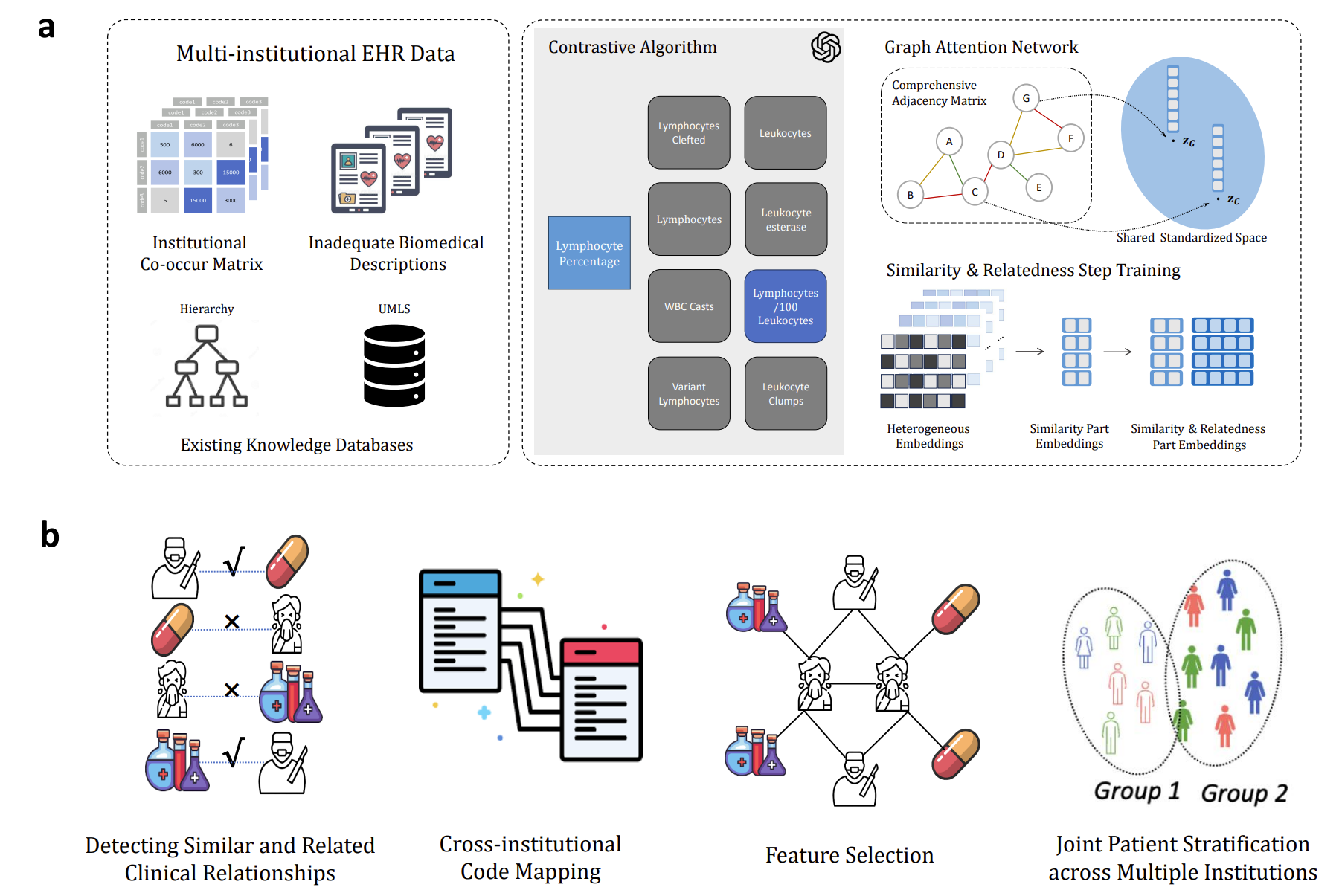
GAME addresses data heterogeneity and privacy constraints in multi-center EHR collaborations by combining graph attention networks and federated learning without raw-data sharing. It integrates information across institutional knowledge graphs, cross-site language-model relations, and a shared contrastive semantic space with hard-negative sampling. Across seven institutions and two languages, GAME demonstrated strong top-1 alignment performance and transferability in Alzheimer’s and suicide-risk studies.
- 5
Semi-supervised triply robust inductive transfer learning
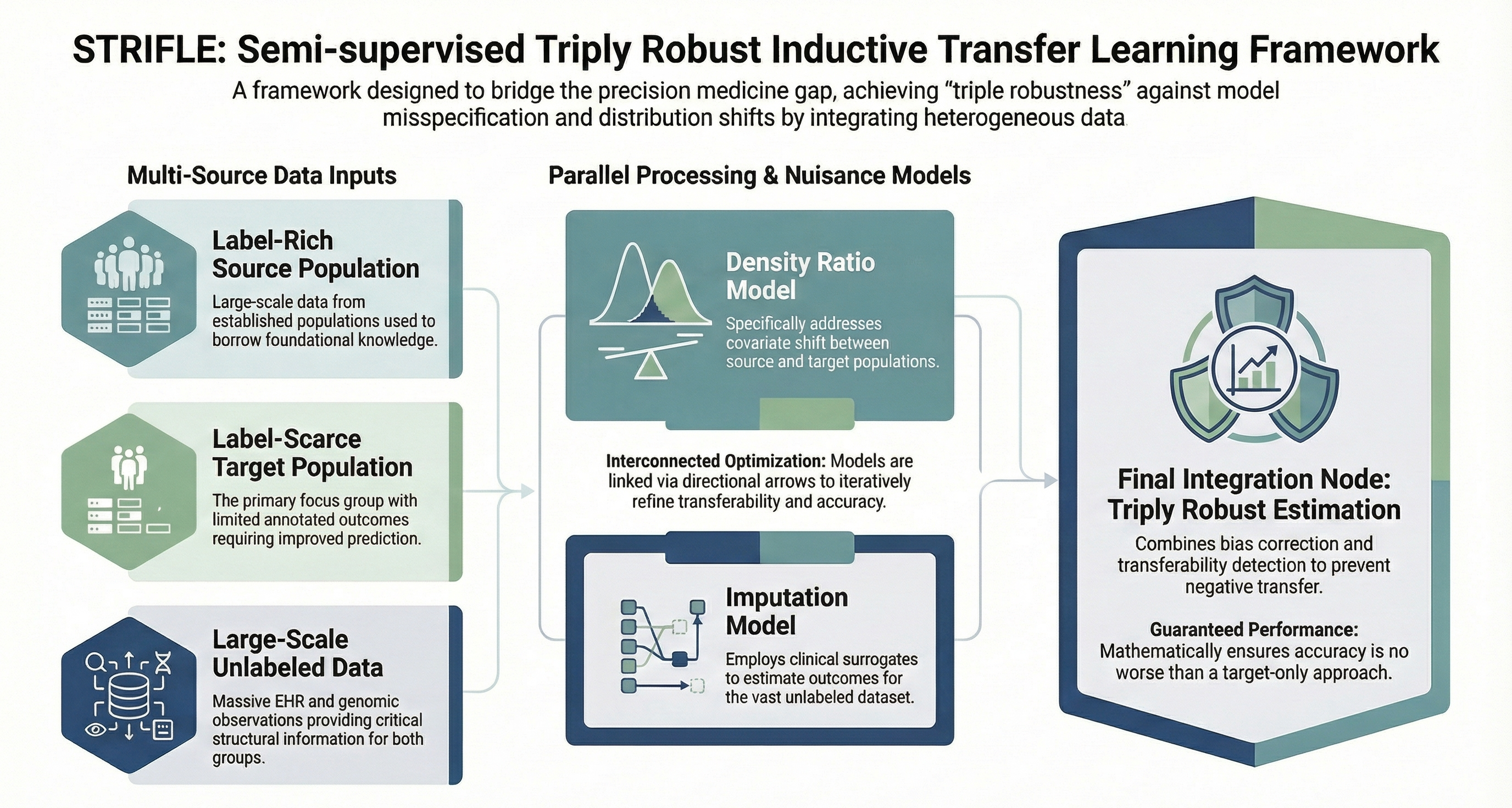
STRIFLE is designed for label-scarce medical settings with high-dimensional covariate shift. It jointly uses source-labeled, target-sparse-labeled, and target-unlabeled data through density-ratio and imputation nuisance models to achieve triple robustness. The method improves efficiency even under nuisance-model misspecification and offers performance guarantees relative to target-only estimation. It is demonstrated in cross-population polygenic risk prediction for Type II diabetes.
- 6
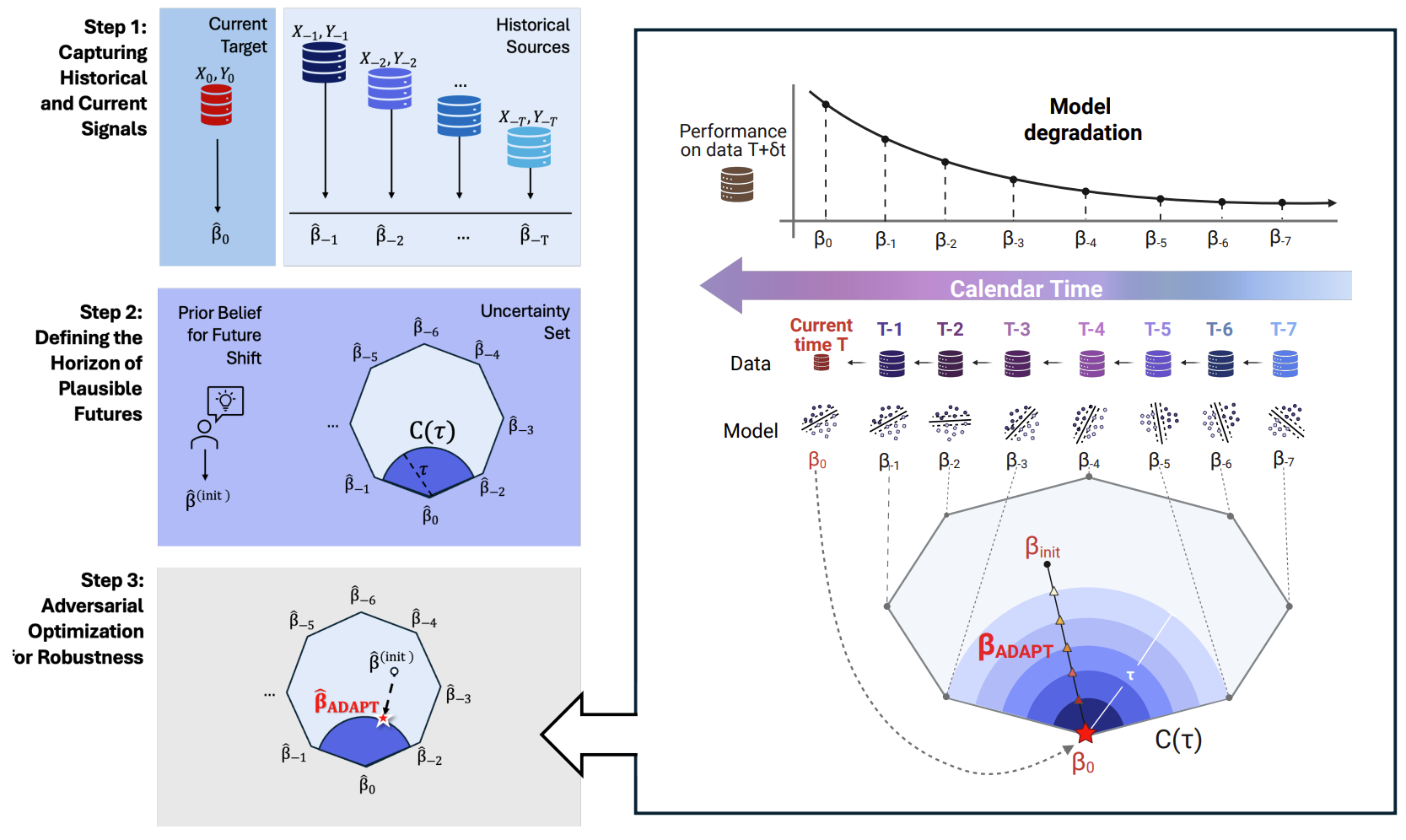
Adversarial Drift-Aware Predictive Transfer: Toward Durable Clinical AI
ADAPT addresses post-deployment performance decay caused by temporal drift, including coding transitions and pandemic shocks. It formulates training as adversarial optimization over an uncertainty set of plausible future distributions inferred from historical estimators and limited current data. By optimizing worst-case future performance, ADAPT balances short-term accuracy and long-term robustness. Longitudinal suicide-risk studies show improved stability without frequent retraining or large new labeling efforts.